 treedata (G, root, attrs, ident, children) Returns data in tree format that is suitable for JSON serialization and use in Javascript documents. Share answered at 3:20 maxkfranz 11. Just point Cytoscape.js to the particular parts of the JSON you want to use (when you call cytoscape ( myOptions ). cytoscapegraph (data, attrs, name, ident) Create a NetworkX graph from a dictionary in cytoscape JSON format. 1 Cytoscape desktop erroneously names the exported file with the extension. The query is relatively complicated since I have to re-map `_key` in the vertex objects to `id`, and the `_from` and `_to` attributes of the edges to `target` and `to`. cytoscapedata (G, attrs, name, ident) Returns data in Cytoscape JSON format (cyjs). This is a current limitation in this feature. Also, Cytoscape desktop can export both Styles and networks, but cannot read Cytoscape.js CSS-like styles because it is VERY different from Styles for Cytoscape desktop. Instead, Cytoscape Desktop handle those as two separate files. 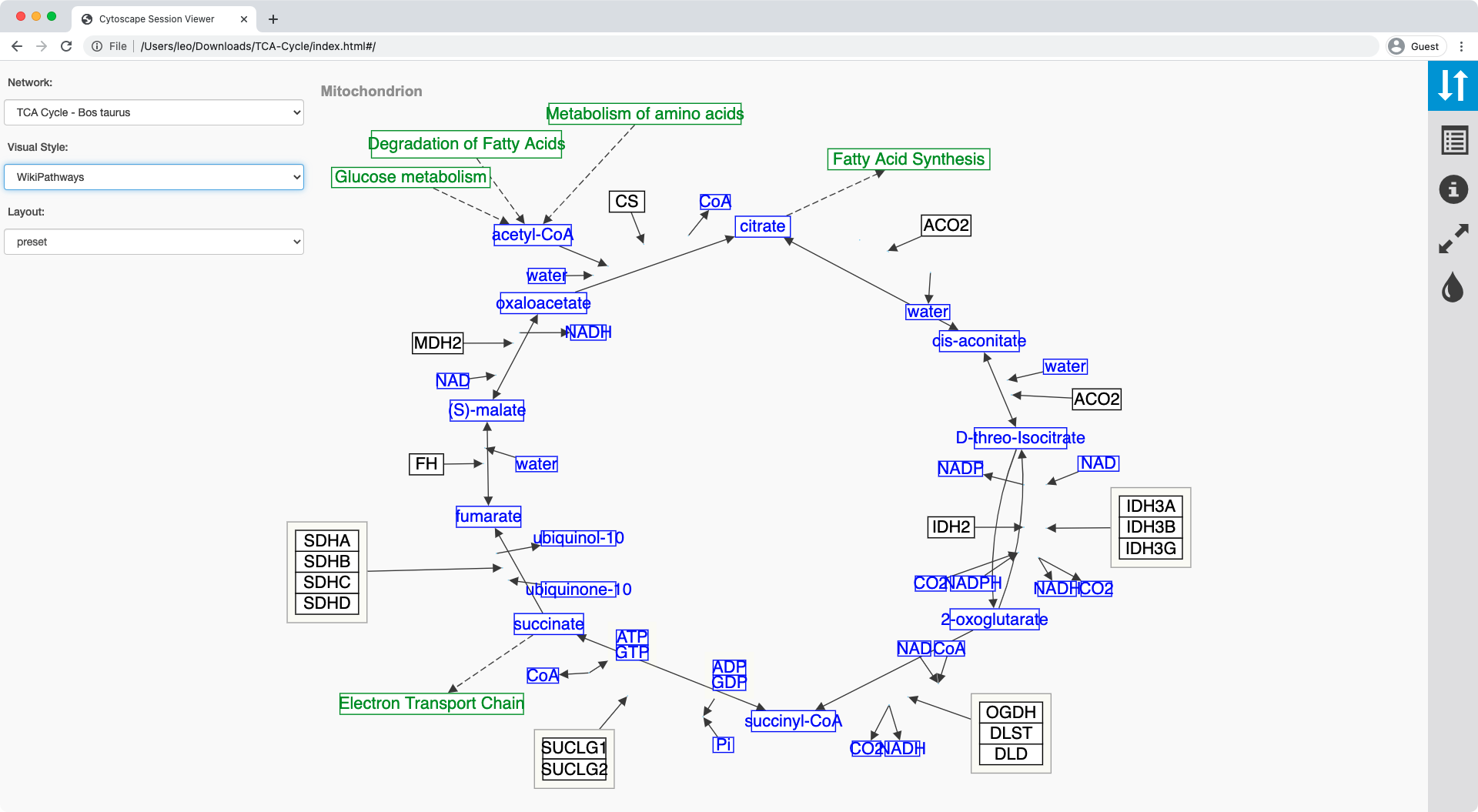
Implementing JSONResult SampleJSONResult is responsible for providing our command’s JSON output. JSON Cytoscape CX The SIF format specifies nodes and interactions only, while other formats store additional information about network layout and allow network data exchange with a variety of other network programs and data sources. The cy.json() exports both Style and network topology in a single JSON. Thus, we transform the original data set by using this Python Notebook. This sample app implements JSONResult with a class called SampleJSONResult and then returns SampleJSONResult from the task ReturnJSONTask. The Cytoscape JSlibrary expects the data to be stored in a JSONfile with a predefined structure. Is this the wrong aproach? or are there more clever ones? This is done via returning implementations of the .JSONResult interface. Which should produce mostly the cytoscape.js format. Require("internal").print(db._query(`LET g = (FOR v, e IN 1.3 OUTBOUND "circles/A" GRAPH "traversalGraph" LET vx = MERGE(v, `).toArray()) I'm using the simple document graph from the arangoDB examples:Īnd using this tiny js scriptlet to get the json file: Simply specify your elements and layout options in the same JSON format as you would use to initialise Cytoscape. 
I'd now like to browse the result with cytoscape and I'm searching for the most elegant way to do this. I've got an ArangoDB database, which can easily format json objects as the result of graph queries. And its not all clear to me whether this is also usefull to read graphs into cytoscape. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
